
By Emily S Finn, Yale University
Each of us is unique, with our own strengths, weaknesses and idiosyncrasies. While this is a truism everyone grasps intuitively, it’s been difficult to determine if and how this individuality is reflected in brain activity.
To investigate, my colleagues and I looked at brain images from volunteers scanned using functional magnetic resonance imaging, or fMRI. This technique measures neural activity via blood flow in the brain while people are awake and mentally active. We calculated a “functional connectivity profile” for each person based on their individual patterns of synchronized activity between different parts of the brain.
In fact, it turns out that the ebb and flow of brain activity is like a fingerprint: each person has their own signature pattern, according to our study just published in the journal Nature Neuroscience. Using only their connectivity profiles, we could identify individuals from a group. Based purely on these profiles, we could also predict how people would perform on one type of intelligence test.
Trading the forest for the trees
fMRI is the best tool we have to study what goes on in a living, thinking human brain in a safe and noninvasive way. And yet fMRI data is notoriously noisy – lots of things influence the signal at any given time, and only some of them are related to the actual brain activity that we care about.
This is why, traditionally, fMRI studies average together data from many different people: the idea is that by finding common patterns of brain activity, we can get rid of much of the noise and end up with something closer to the “true” signal. Essentially, we blend all the individuals’ signals to get one version that’s representative of the whole population.
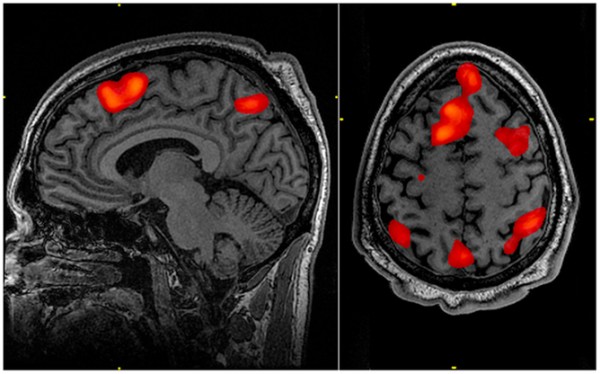
But you don’t need to be a brain scientist to recognize that everyone is different; this averaging probably obscures interesting activity patterns that are idiosyncratic to each person. And for fMRI to be practically useful – in medicine, for example – we’d need to get meaningful information based on a scan from a single person.
We set out to prove that analyzing fMRI data from individual people is indeed possible, by showing that these idiosyncratic activity patterns are reliable enough to identify individuals from a large group.
Analyzing individual scans
We used data from the Human Connectome Project (HCP), a major research effort to collect brain-imaging data along with behavioral, demographic and genetic information from a large number of healthy people. So far, data from 500 people have been released, and there are plans to collect 1,200 in total. All the data are made publicly available, so researchers anywhere can download it, analyze it in different ways, and mine it for interesting insights.
We looked at data from the first 126 participants in the HCP. Each person was scanned six different times. During two of the scans, people were simply resting, letting their minds wander. During the other four scans, they worked on some type of cognitive task: trying to hold items in mind in a test of working memory, listening to a story, solving math problems, looking at emotional faces or moving different parts of their body.
To analyze the fMRI data for each individual participant, we first divided the whole brain into 268 separate regions. While it’s an open question just how many different functional regions there are in the brain, previous work of ours has shown that using between 200 and 300 regions lets us detect subtle effects, while still keeping things manageable in terms of the time and computing power it takes to run the analyses.
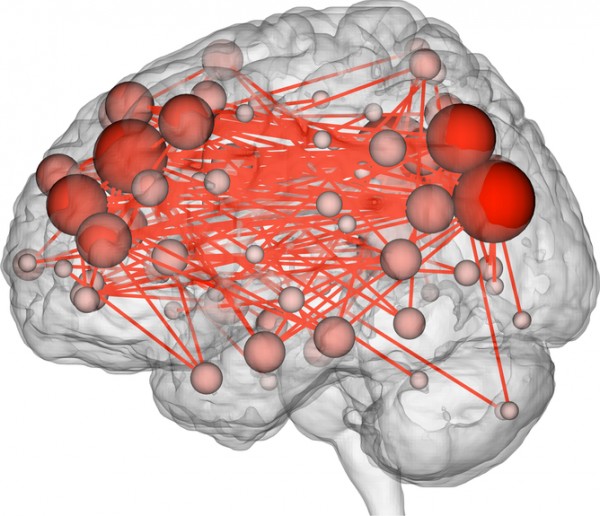
For each pair of regions, we calculated the strength of the functional connection between them. To understand what a “functional connection” is, think of two musicians playing at the same time: rather than measuring how loudly each musician is playing, we measure how synchronized their playing is. It’s not about overall levels of activity in any single brain region, but rather how pairs of regions tend to increase and decrease their activity in tandem. We calculated this measure of synchrony for every pair of regions across a brain. For each person we had a functional connectivity profile for each of the six scans they underwent.
We wanted to see if connectivity profiles could act like fingerprints. So we took a single profile from one scan session – say, the working memory session – and compared it to all 126 profiles for a different scan session, say the one at rest. Based on the numerical profiles, we figured out which other profile was its closest match. Would we be able to match up the participant’s working memory and at-rest scans? That is, would an individual’s brain “look the same” no matter what task it was doing?
The majority of the time, the identity we had predicted was indeed the correct one: we were able to identify people with up to 99% accuracy. The accuracy ranged from 64% to 99%, depending the specific pair of scan sessions. If we were just randomly guessing, we would expect to choose the right identity less than 1% of the time, so this was a very significant result.
Predicting fluid intelligence
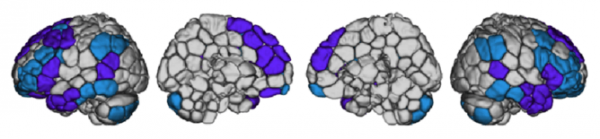
Certain connections were most distinguishing of individuals – namely, those between the brain’s prefrontal lobe (just behind the forehead) and parietal lobe (farther back on top of the head). These areas evolved most recently, and neuroscientists have long known that they are crucial for sophisticated functions like attention, memory and language.
We discovered that these connections could also predict how people would perform on a test of fluid intelligence, or on-the-spot reasoning ability. Fluid intelligence is the ability to see patterns and solve reasoning problems.
While the predictions of fluid intelligence were overall more accurate than not, there was still a fair amount of error – the model overpredicted some people’s scores and underpredicted others’ – so we certainly wouldn’t advocate giving someone a brain scan instead of an IQ test or other traditional assessment.
In the first part of our study, we found that people always look most like themselves, regardless of what they’re doing. In other words, the same brain doing two different tasks always looks more similar than two different brains doing the same task. And in the second part of our study, we saw that these connectivity profiles correspond to highly complex cognitive attributes.
Why does this matter? After all, we don’t need to put someone in an MRI scanner to know who they are – we can tell that by looking at them. The importance of this finding is that these connectivity profiles could potentially give us information about people that is harder to tell just by looking.
For example, they could help predict who is at risk for developing a disease. Maybe there’s something in the individual patterns of strong and weak brain connections that reveals how susceptible someone is to different neurological or mental illnesses, such as schizophrenia, depression or Alzheimer’s disease. If we collect fMRI images from people while they are still healthy, and then follow them over time to see who goes on to become ill, perhaps we can build a model relating parts of the connectivity profile to future health. Then we could apply this model to a brand-new person’s profile to predict their likelihood of getting sick. This could be a way to target and treat high-risk people early on, in hopes that intervening early will improve their outcomes.
Ultimately, we hope these profiles could someday be used in personalized medicine, a way to customize interventions and therapies for people based on their individual biology.
But there are still many open questions. For example, we tested identification between scans separated by a few days, but how stable are connectivity profiles over a period of months or years? Can they change as a function of aging, illness, cognitive training or some other process? What other behavioral traits are reflected in patterns of brain connectivity? While there is much work to be done, my colleagues and I believe that these results provide an exciting foundation for future research.
![]()
Emily S Finn, PhD Candidate in Neuroscience, Yale University
This article was originally published on The Conversation. Read the original article.











